One intensively investigated example is antibody-based treatment targeting tumor necrosis factor alpha (TNF), which has been successfully used to treat a range of autoimmune/inflammatory diseases such as rheumatoid arthritis (RA) and inflammatory bowel disease (IBD). For both of these diseases, most patients respond very well to anti-TNF therapy, whereas a significant minority show no response at all. A recent study by Prasad et al. used Olink-based plasma proteomics to look for protein profiles that could potentially predict which RA patients would or would not respond to six months of anti-TNF treatment. As summarized in the figure below, plasma proteomics identified a machine learning-derived identifier composed of 17 proteins that can predict treatment responders with high accuracy. Additionally, a principal component analysis (PCA) of the baseline samples clustered patients into two previously unknown RA endotypes, which could further help refine and guide future treatment.
Predicting drug responders in rheumatoid arthritis (RA)
Background
- Anti-TNF therapy is expensive – no benefit to 1/3 of RA patients.
Method
- Plasma from 144 RA patients on anti-TNF therapy interrogated with four Olink® Target 96 panels.
- Baseline samples examined by PCA analysis – machine learning applied to samples grouped by response/non-response after 6 months – can protein biomarker profiles predict responders?
Results
- PCA analysis – 2 clusters of RA patients with clinical differences
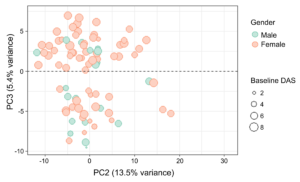
Prasad et al. (2022) PLoS Computational Biology, DOI: 10.1371/journal.pcbi.1010204
- Machine learning identifies a 17-protein classifier to identify responders (AUC=0.88).
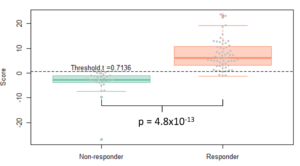
Prasad et al. (2022) PLoS Computational Biology, DOI: 10.1371/journal.pcbi.1010204
Conclusions
- Identified novel endotypes (gender & disease severity) for RA that could aid future treatment development.
- Predictive classifier for responders helps optimize treatment selection, reduce costs and improve quality of life for non-responders.



